Epigenomic: CRISPRi and CRISPRa
What is dCas9?
Cas9 from Streptococcus pyogenes is the most frequently used cas endonuclease in CRISPR. The dCas9 (deficient Cas9 or Dead cas9) is the mutant of spCas9. It lacks the ability to cleave double-stranded DNA but is still capable of cleaving only one strand of the genome DNA (so-called “nickase”). Like Cas9, dCas9 can still be localized to the desired genome site guided by specific RNAs (gRNA), thus binding to the targeted genomic DNA sites..
What is the application for dCas9?
The dCas9 protein binds to a specific genomic location under the guidance of a target site-specific gRNA. When coupled with a protein, the resulting “dCas9-Protein” fusion can modify the genome sequence by preventing the cell’s transcription machinery from accessing the binding region, activating promoters, or inducing CRISPR interference (CRISPRi) or activation (CRISPRa). The activation domains recruited to the specific target site by the dCas9 protein and gRNA, do not affect the expression of other genes in the genome. This makes it an excellent tool for epigenomics analysis. Refer to the illustration scheme below for CRISPR-based gene regulation.
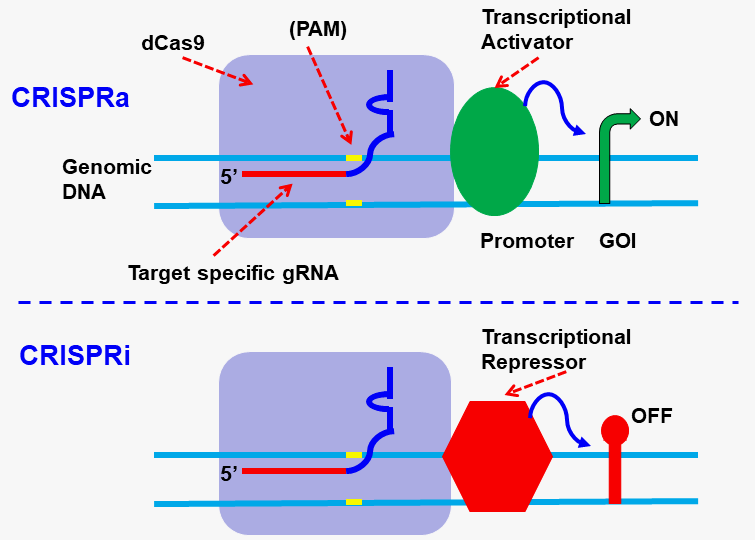
Gentarget’s dCas9 products?
Gentarget Inc generated the dCas9 expression construct by making two point mutation in spCas9’s endonuclease domains (RuvC and HNH), as D10A and H840A, resulting in its deactivation. The dCas9 expression is in-framed with an enhanced NLS (nuclear localization signal), and a HA tag (to purify or detect the expressed dCas9 when desired) .
The dCas9 protein binds to a specific genomic location under the guidance of a target site-specific gRNA. When coupled with a protein, the resulting “dCas9-Domain” fusion can activate or repress the targeted gene expression depend upon the fusion protein property. It is a great tool for Epigenomic assays, such as gRNA library screening for epigenomic activation or repression.
See the illustration scheme below of CRISPR based gene regulation.
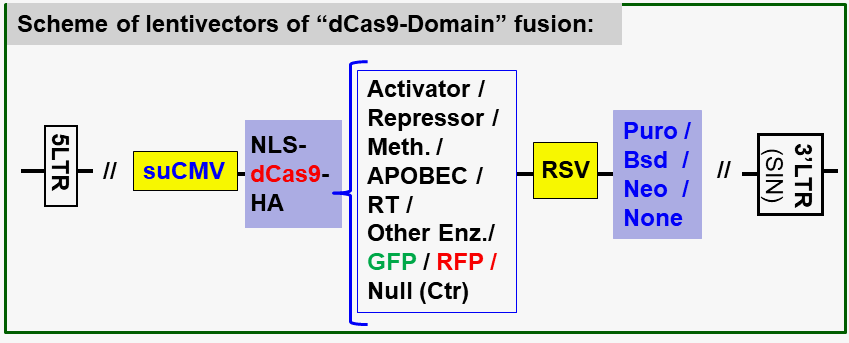
Please look into each product page below, or click the Product Manual for details.